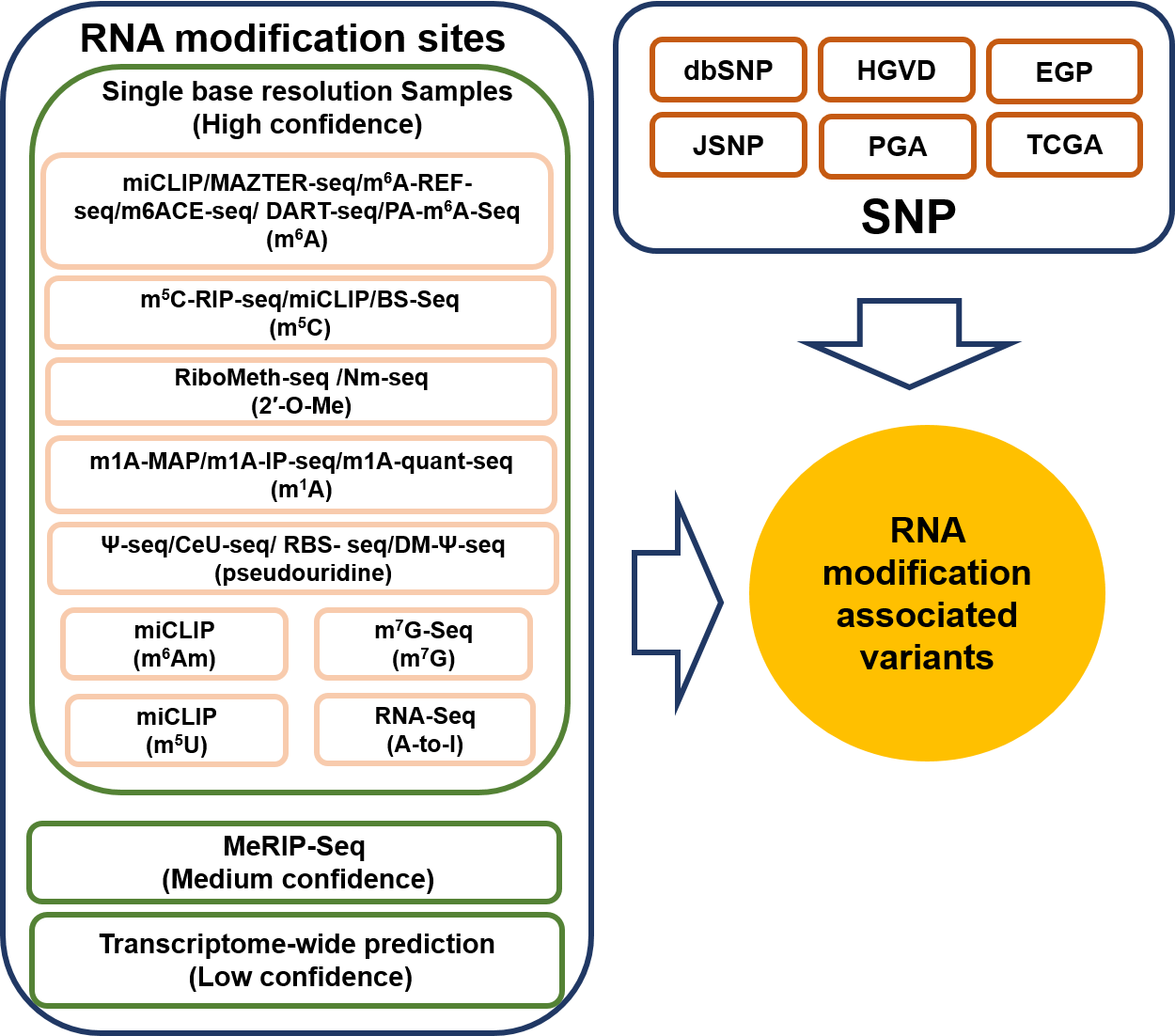
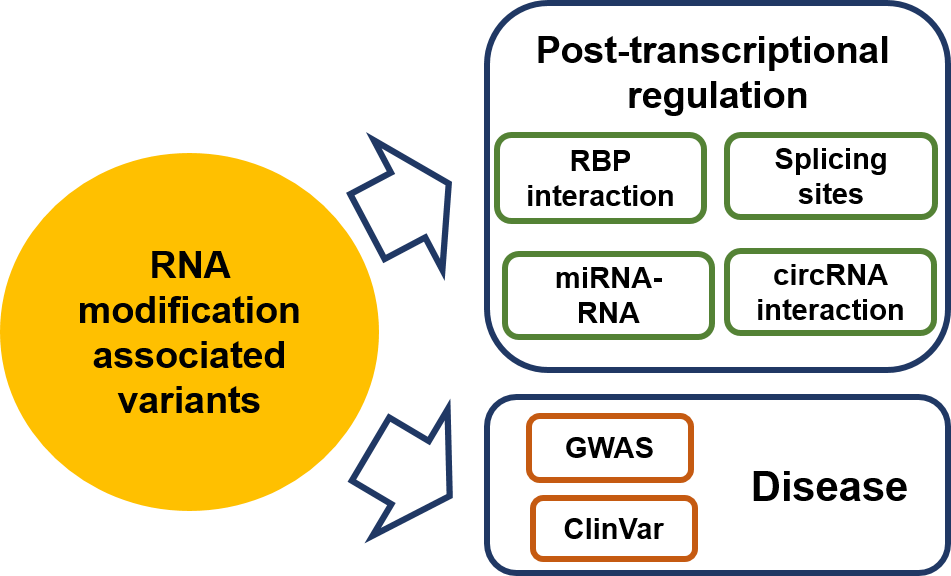
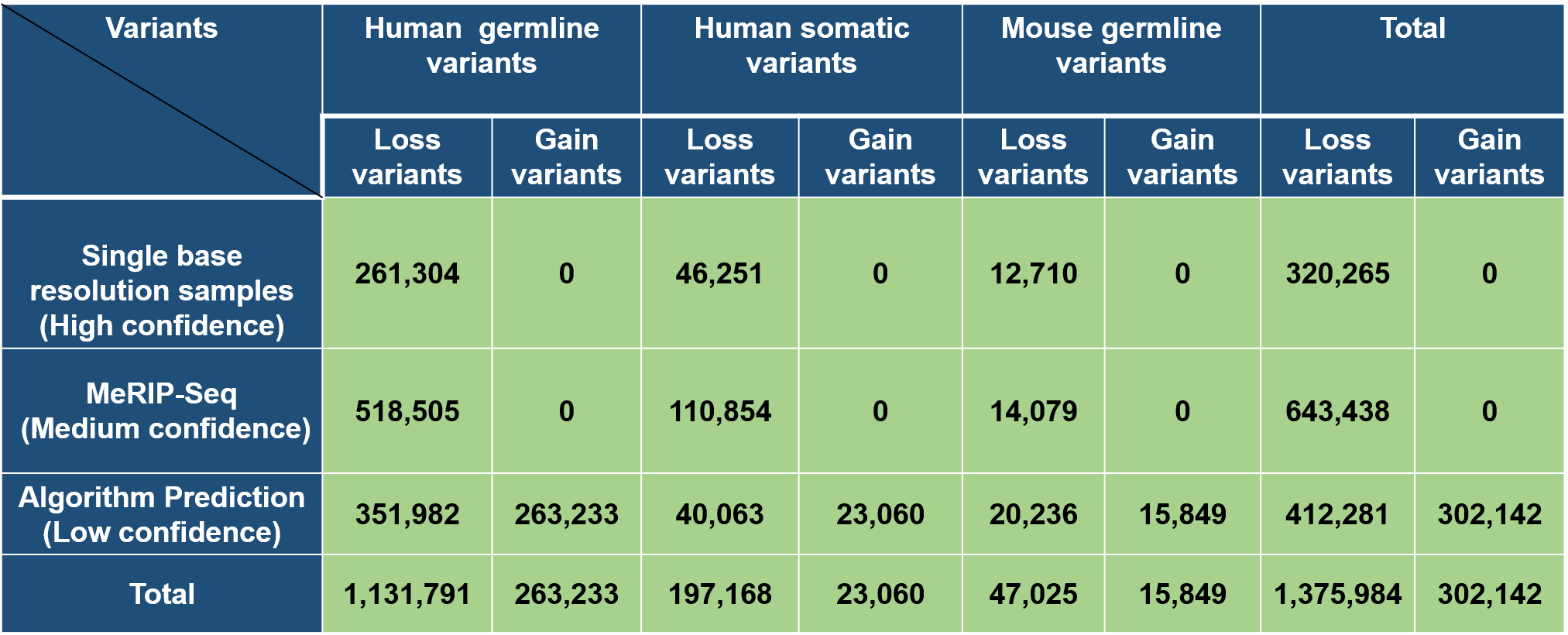
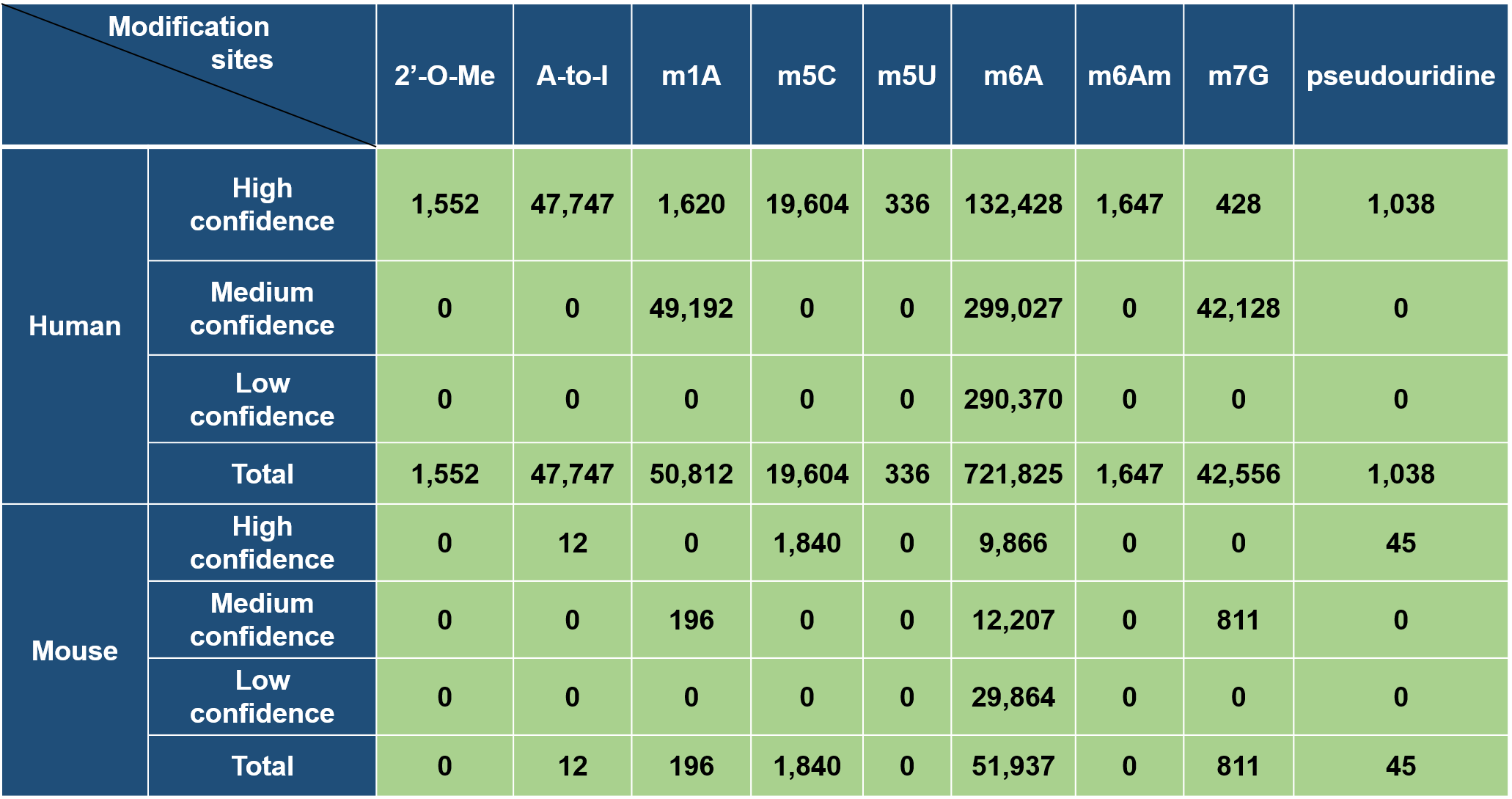
According to different levels of RNA modifications sites confidence, we extracted the corresponding RNA modifications associated variants as follows: 1. For RNA modifications sites with high confidence level, we retained the variants that located nearby the RNA modifications sites and then looked for the variants that destroy modification motif of the modification sites. 2. For RNA modifications sites with medium confidence level, the RNA modification associated variants were derived from the intersection between the variants and RNA modification sites generated from MeRIP-Seq experiments and m6A-Seal-Seq experiments. Then, we applied our prediction models to find the variants nearby the RNA modification sites that change the sequence features essential for RNA modification. 3. For RNA modification sites low confidence level, a genome-wide prediction based on deep convolutional neural network was performed for the sequences around all the collected variants from dbSNP, HGVD, TCGA, ICGC and COSMIC to find the variants that cause potential gain or loss of RNA modification sites.